


Crucifer Genome Initiative
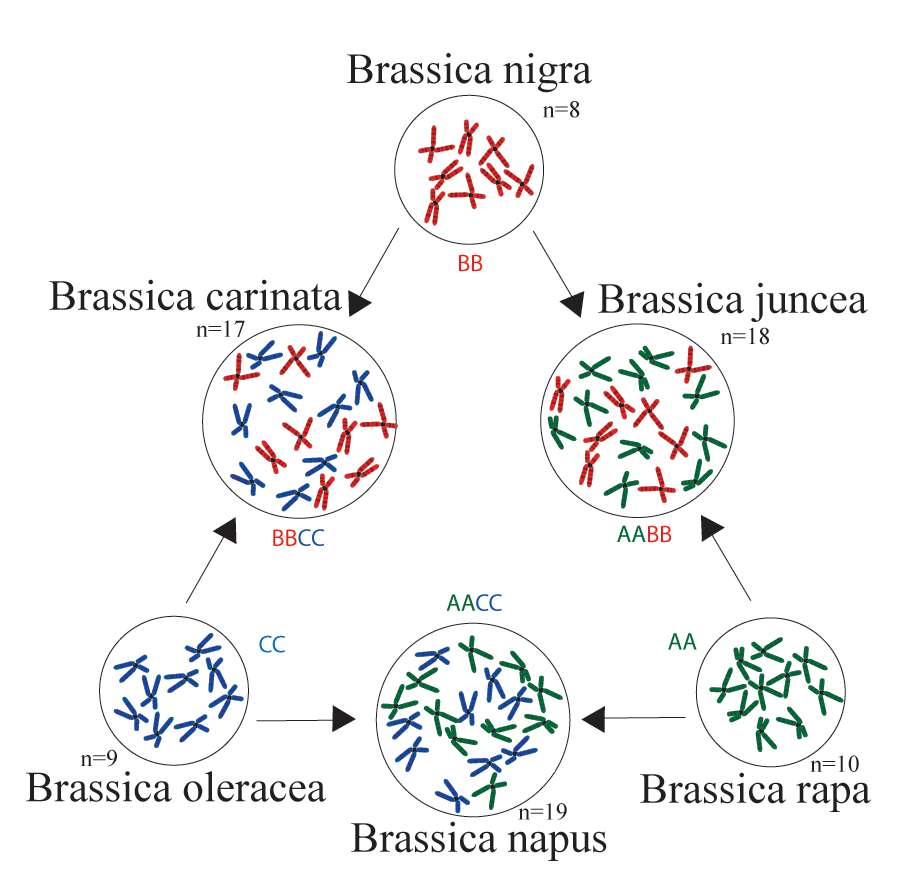
The Crucifer Genome Initiative (CGI) is a collaboration between Agriculture and Agri-Food Canada Saskatoon Research and Development Centre and the Global Institute for Food Security, Saskatoon (https://www.gifs.ca) to provide access to a growing collection of genome sequences and associated resources that are being developed for a number of species of the Brassicaceae family. The group in Saskatoon initially focused on Brassica species of U’s triangle (U, 1935) contributing to international efforts to sequence Brassica rapa (Wang et al, 2009) and Brassica napus (Chalhoub et al, 2014) and published a genome sequence for Brassica oleracea (Parkin et al, 2014). Interest in establishing new oilseed species for the Canadian Prairies led to the generation of a genome sequence for Camelina sativa (Kagale et al, 2014) and short read based genome sequences have also been developed for Brassica juncea, Brassica carinata, Sinapis alba and a Canadian spring type B. napus line. With the adoption of oxford nanopore technology’s long read sequencing, our ability to rapidly generate high quality reference genomes at reasonable cost has allowed us to add further genomes to our collections. Initial proof of concept for ONT derived genome sequences of polyploid genomes was carried out for Brassica nigra (Perumal et al, submitted). Collaborative efforts are now underway to sequence multiple Camelina and Lepidium species and further genotypes of B. napus. Please refer to the Contact page where the numerous sources of funding and our colleagues who have supported the development of these resources are listed.